Gene Test Can Predict Risk of Medications Causing Liver Injury
Research By: Takanori Takebe, MD
Post Date: September 7, 2020 | Publish Date: Sept. 7, 2020
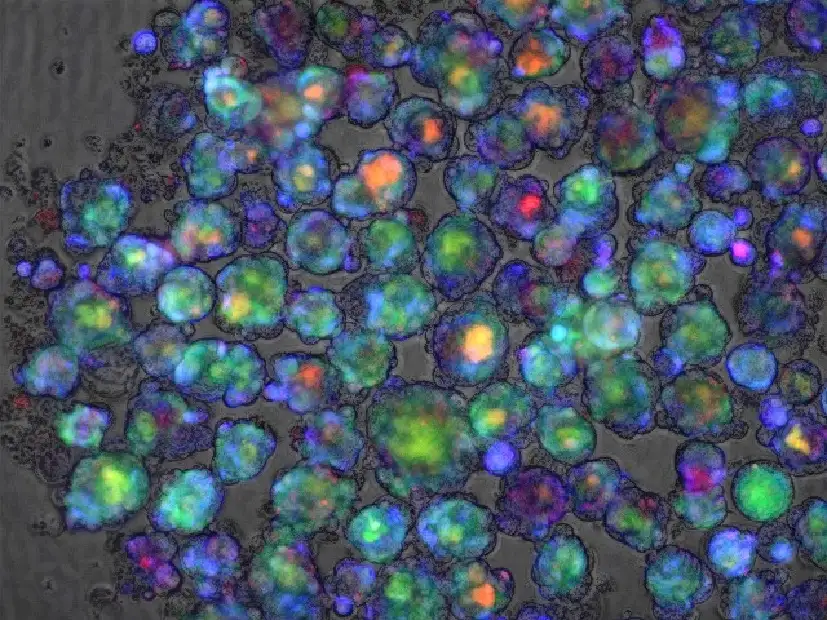
Organoid-Related Discovery May Also Help Drug Makers Screen Potential Pharmaceuticals
Scientists who were working on a way to determine the viability of batches of tiny liver organoids have discovered a testing method that may have far broader implications.
Their study published Sept. 7, 2020, in Nature Medicine, reports identifying a “polygenic risk score” that shows when a drug, be it an approved medication or an experimental one, poses a risk of drug-induced liver injury (DILI).
The work was conducted by consortium of scientists from Cincinnati Children’s, Tokyo Medical and Dental University, Takeda Pharmaceutical Co. in Japan, and several other research centers in Japan, Europe and the US. The findings take a large step toward solving a problem that has frustrated drug developers for years.
“So far we have had no reliable way of determining in advance whether a medication that usually works well in most people might cause liver injury among a few,” says Jorge Bezerra, MD, Director, Division of Gastroenterology, Hepatology and Nutrition at Cincinnati Children’s.
“That has caused a number of promising medications to fail during clinical trials, and in rare cases, also can cause serious injury from approved medications. If we could predict which individuals would be most at-risk, we could prescribe more medications with more confidence,” says Bezerra, who was not involved with the study.
Now that reliable test might be just around the corner.
“Our genetic score will potentially benefit people directly as a consumer diagnostic-like application, such as 23andMe and others. People could take the genetic test and know their risk of developing DILI,” says corresponding author Takanori Takebe, MD, an organoid expert at Cincinnati Children’s who has been studying ways to grow liver “buds” for large-scale use in research.
The team developed the risk score by re-analyzing hundreds of genome-wide association studies (GWAS) that had identified a long list of gene variants that might indicate a likelihood of a poor reaction in the liver to various compounds. By combining the data and applying several mathematical weighting methods, the team found a formula that appears to work.
- The risk score takes more than 20,000 gene variants into account.
- The team confirmed the score’s prediction power in cell culture, in organoid tissue and by using patient genomic data already on file.
- The score was valid in tests involving more than a dozen medications: cyclosporine, bosentan, troglitazone, diclofenac, flutamide, ketoconazole, carbamazepine, amoxicillin–clavulanate, methapyrilene. tacrine, acetaminophen and tolcapone.
- The test works for different types of drugs because the score focuses on a set of common mechanisms involved in how the liver metabolizes a drug, including oxidant stress pathways in liver cells and endoplasmic reticulum (ER) stress—a disruption of cell function that happens when proteins cannot fold properly.
How can a risk score help?
For clinicians, this would allow them to run a quick genetic test to identify patients at higher risk of liver injury before prescribing medications. The results might prompt a doctor to change the dosage, order more frequent follow-up tests to catch early signs of liver damage, or switch medications entirely.
For drug research, the test could help exclude people of high risk of liver injury from a clinical trial so that the benefits of a medication can be more accurately assessed.
Liver toxicity has caused a number of drug failures over the years. Takebe says both patients and the drug maker were disappointed when a potential diabetes treatment called fasigliam was withdrawn in 2014 during phase 3 clinical trials. Some of the participants (at a rate equivalent to about 1 in 10,000) experienced elevated enzyme levels that suggested potential liver injury.
While such risks may appear low, at the time there was no way to predict which people would develop DILI, making the drug unacceptably dangerous. But the new polygenic risk score would make it possible to produce liver organoids that exhibit key risk variants to determine if a drug is harmful before people ever take it.
What’s next?
Takebe and colleagues demonstrated how to produce liver buds on a mass scale in 2017 in a study published in Cell Reports. The team has improved upon the process since, reporting success in 2019 in Cell Metabolism at engineering liver organoids that model disease.
However, more research involving a more-diverse population is needed to confirm the initial findings and to scale up a DILI screening test for potentially widespread use, Takebe says.
About the study
In addition to Cincinnati Children’s, collaborators on this study included experts from Yokohama City University, the T-CiRA Joint Program, Takeda Pharmaceutical Company, and the Tokyo Medical and Dental University in Japan; Nottingham and Newcastle Universities in the United Kingdom; the Icahn School of Medicine at Mount Sinai in New York, and the University of North Carolina,
Funding sources primarily include Takeda Pharmaceutical Company via the T-CiRA program (Director, Shinya Yamanaka).Sources also include the National Institutes of Health (UG3 DK119982), , the New York Stem Cell Foundation, Cincinnati Children’s Research Foundation, the Dr. Ralph and Marian Falk Medical Research Trust Awards Program, the Takeda Science Foundation, the Mitsubishi Foundation award, and AMED JP19fk0210037, JP19bm0704025, JP19fk0210060, JP19bm0404045 and JSPS JP18H02800,19K22416.
| Original title: | Polygenic architecture informs potential vulnerability to drug-induced liver injury |
| Published in: | Nature Medicine |
| Publish date: | Sept. 7, 2020 |
Research By