Viruses Play Critical Role in Evolution and Survival of the Species
Research By: Satoshi Namekawa, PhD
Post Date: September 7, 2020 | Publish Date: Sept. 7, 2020

“What we learn from our study is that, in general, viruses have major roles in driving evolution. In the long-term, viruses have positive impacts to our genome and shape evolution.”
–Satoshi Namekawa, PhD
As the world scrambles to control the growing COVID-19 coronavirus pandemic, new research in Nature Structural & Molecular Biology shows viruses also play a key evolutionary role in mammals’ ability to reproduce and survive.
Scientists in the Cincinnati Children’s Perinatal Institute and at Azabu University in Japan obtained their data by studying laboratory mice and human germline cells.
In two separate papers appearing in the same edition of the journal, they reveal two distinct and fundamental processes underlying germline transcriptomes. They also show that species-specific transcriptomes are fine-tuned by endogenous retroviruses in the mammalian germline.
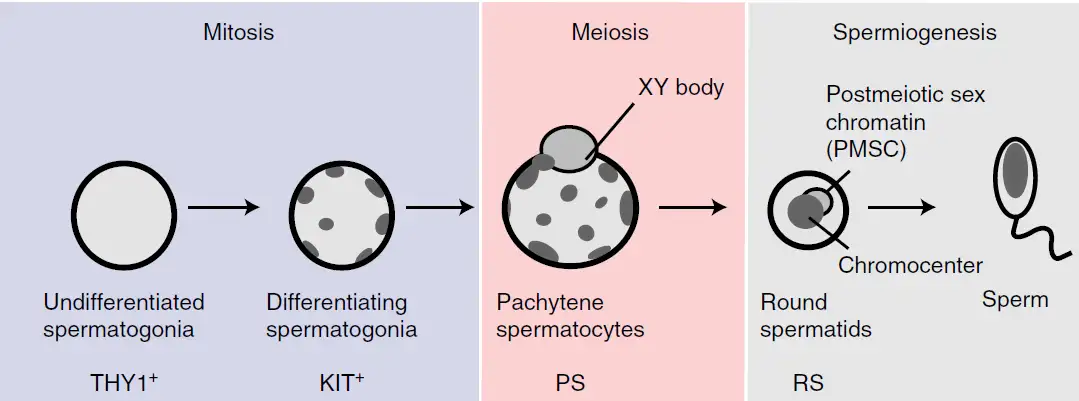
Germline transcriptomes include all the messenger RNA in germline cells, which contain either the male or female half of chromosomes passed on as inherited genetic material to offspring when species mate. This means that germline transcriptomes define the unique character of sperm and egg to prepare for the next generation of life.
Although the studies are separate they complement one another, according to Satoshi Namekawa, PhD, principal investigator on both papers and a scientist in the Division of Reproductive Science at Cincinnati Children’s.
“One paper, Maezawa and Sakashita et, al., explores super-enhancers, which are robust and evolutionally conserved gene regulatory elements in the genome. They fuel a tightly regulated burst of essential germline genes as sperm start to form,” Namekawa said.
“The second study, Sakashita et, al., involves endogenous retroviruses that act as another type of enhancer – gene regulatory elements in the genome – to drive expression of newly evolved genes. This helps fine tune species-specific transcriptomes in mammals like humans, mice, and so on.
Clinical Relevance
Together the studies have significant potential ramifications for clinical practice, according to study authors, who include a multi-disciplinary mix of developmental biologists, bioinformaticians and immuno-biologists. Dysregulation of gene expression in the formation of male sperm is closely associated with male infertility and birth defects.
Viruses, especially endogenous retroviruses (ERVs) that are an inherent part of mammalian biology, can dramatically influence gene expression, investigators report. ERVs are molecular remnants of retroviruses that infect the body and over time incorporate into the genome.
“What we learn from our study is that, in general, viruses have major roles in driving evolution,” Namekawa explained. “In the long-term, viruses have positive impacts to our genome and shape evolution.”
Super-Enhancer Switch
The study, Maezawa and Sakashita et, al., combined biological testing of mouse models and human germline cells with computational biology, including genome-wide profiling of gene regulatory elements in germline cells.
Those tests revealed that the the genome-wide reorganization of super-enhancers drives bursts of germline gene expression after germ cells enter meiosis, a specialized form of cell division that produces the haploid genome of germ cells.
The study further demonstrates the molecular process through whichsuper-enhancer switching takes place in germ cells. Super-enhancers are regulated by two molecules that act as gene-burst control switches – the transcription factor A-MYB and SCML2, a critical silencing protein in sperm formation.
TEs and Jumping Genes
Endogenous retroviruses are a group of transposable elements (TEs), mobile genetic elements that account for approximately 40–50 percent of a given mammalian genome. Also referred to as “jumping genes,” TEs have long been considered genetic threats because transposition can be harmful if, for example, the process disrupts protein-coding genes.
Building on findings from the 1950s that TEs can function as genetic regulatory elements, Namekawa and his collaborators (Sakashita et, al.) produced data showing that ERV-driven mechanisms help fine tune species-specific transcriptomes.
About the Study
Funding support for the ERV study came in part from: the National Institutes of Health (R01 GM122776, DP2 GM119134); a Lalor Foundation Postdoctoral Fellowship; a grant from the Azabu University Research Services Division; Japan’s Ministry of Education, Culture, Sports, Science and Technology (MEXT)-Supported Program for the Private University Research Branding Project (2016–2019); a Grant-in-Aid for Research Activity Start-up (19K21196); the Uehara Memorial Foundation Research Incentive Grant; an Albert J. Ryan Fellowship, and a March of Dimes Prematurity Research Centre Collaborative Grant (#22-FY14-470).
Support for the super-enhancer study came in part from a research project grant by the Azabu University Research Services Division; a grant from Japan’s Ministry of Education, Culture, Sports, Science and Technology and (MEXT)-Supported Program for the Private University Research Branding Project (2016–2019); a Grant-in-Aid for Research Activity Start-up (19K21196) from the Takeda Science Foundation (2019); the Uehara Memorial Foundation Research Incentive Grant (2018); a Lalor Foundation Postdoctoral Fellowship; an Albert J. Ryan Fellowship; and a Cincinnati Children’s Endowed Scholar Award; CpG grant awards and the National Institute of Health (DP2 GM119134, R01GM122776 and GM098605)
This post was written by Nick Miller, Cincinnati Children’s
Read more about how viruses can rewrite parts of the code that makes us human:
Gene Discovery Reveals How Human Pregnancy Evolved Away from Marsupial Pouches
| Original title: | Super-enhancer switching drives a burst in gene expression at the mitosis-to-meiosis transition |
| Published in: | Nature Structural & Molecular Biology |
| Publish date: | Sept. 7, 2020 |
Research By