25 Years Later, Xenbase Still Supporting Scientific Leaps
Research By: Aaron Zorn, PhD
Post Date: December 24, 2025 | Publish Date: Nov. 3, 2025
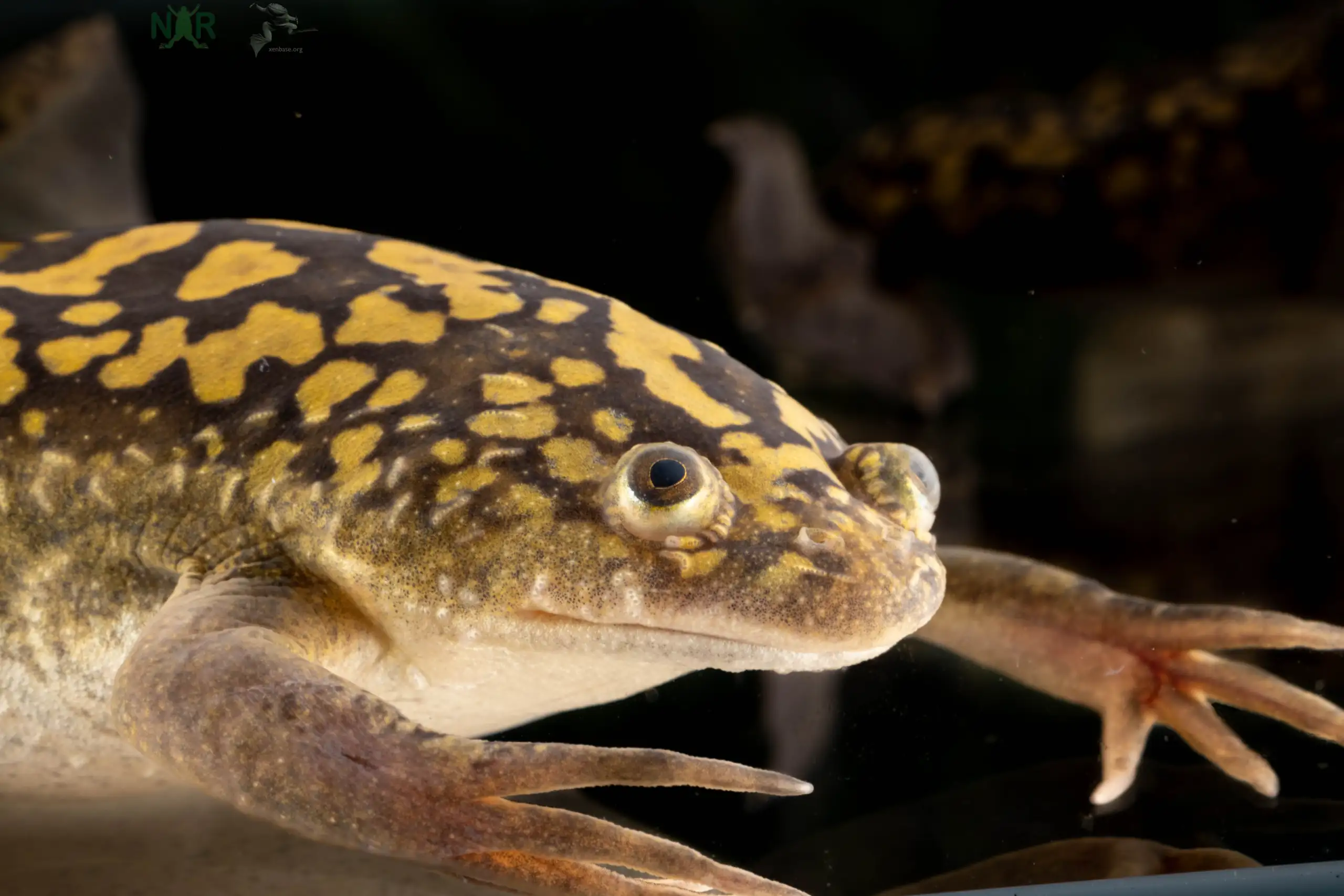

As it shovels in food with a set of stubby finger-like claws, the slimy African clawed frog isn’t likely to win many beauty contests. But in its alter ego form—Xenopus laevis—the tadpoles of this homely amphibian are scientific superheroes.
This frog is one of the most-thoroughly studied creatures on Earth. In the 1940s, it was widely used in pregnancy testing. In 1958, it was the first vertebrate to ever be successfully cloned. In 1992, NASA sent four African clawed frogs into orbit to study their ability to reproduce in zero gravity. In 2016, the first complete genome of Xenopus laevis was sequenced.
With over 90% of the human genes associated with disease having frog orthologs, Xenopus is a powerful model for modeling the developmental origins of health and disease. And few people on Earth are as familiar with Xenopus as Aaron Zorn, PhD.
Since 2008, Zorn has worked with Peter Vize, PhD, of the University of Calgary, as co-directors of Xenbase—the world’s biggest and constantly evolving collection of genomic and developmental data about the African clawed frog. Xenbase has grown to include more than 1,700 registered users worldwide who tap into curated data on gene expression, functional genomics and more.
Earlier this year, this priceless scientific repository celebrated its 25th anniversary. Zorn, Vize and colleagues marked the occasion with an article published in Genetics entitled “Xenbase: 25 years of integrating molecular and biomedical data from Xenopus.”
To this day, Cincinnati Children’s hosts the bio-curation team that processes scientific publications from around the world that involve Xenopus, annotating the frog’s genome and parsing new findings into the database. Curators document which genes are examined, where they are expressed, what antibodies and other reagents were used, and how the experiments influence embryonic development.
Meanwhile, the team in Calgary runs the computational side of the project, including server management, the database, the custom web application and user interfaces. The project has been funded for years by the Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD).
These days, Zorn directs the Division of Developmental Biology at Cincinnati Children’s. He also serves as co-director of the Center for Stem Cell and Organoid Medicine (CuSTOM), which has been a leading force in advancing regenerative medicine.
In recognition of reaching the silver anniversary milestone, we asked Zorn to reflect on the far-reaching impact of Xenbase.
What was the inspiration for Xenbase?
For many years we have relied on animal models to study the basics of developmental biology. Flies, worms, fish, frogs and mice. Those are the main ones used in the scientific community. About 30 years ago, it was recognized that there were thousands of labs publishing thousands of papers using these models. Entire genomes were being sequenced, and there was no way to organize all this material, to curate all this information. We needed a way to make these papers computer readable. There was already a FlyBase and the MGI database in development for mice. So that’s where Xenbase came along for the Xenopus community.
How did you get involved?
I started on this back in 2008, right around when we got our first big NICHD resource grant. Now we have a team of a half-dozen staff scientists who are bio-curators. These are people with PhDs and postdocs who interact with the research community, with the NCBI (National Center for Biotechnology Information) and with these other organism databases. They curate the literature. They make connections to disease models. And they represent us at international meetings.
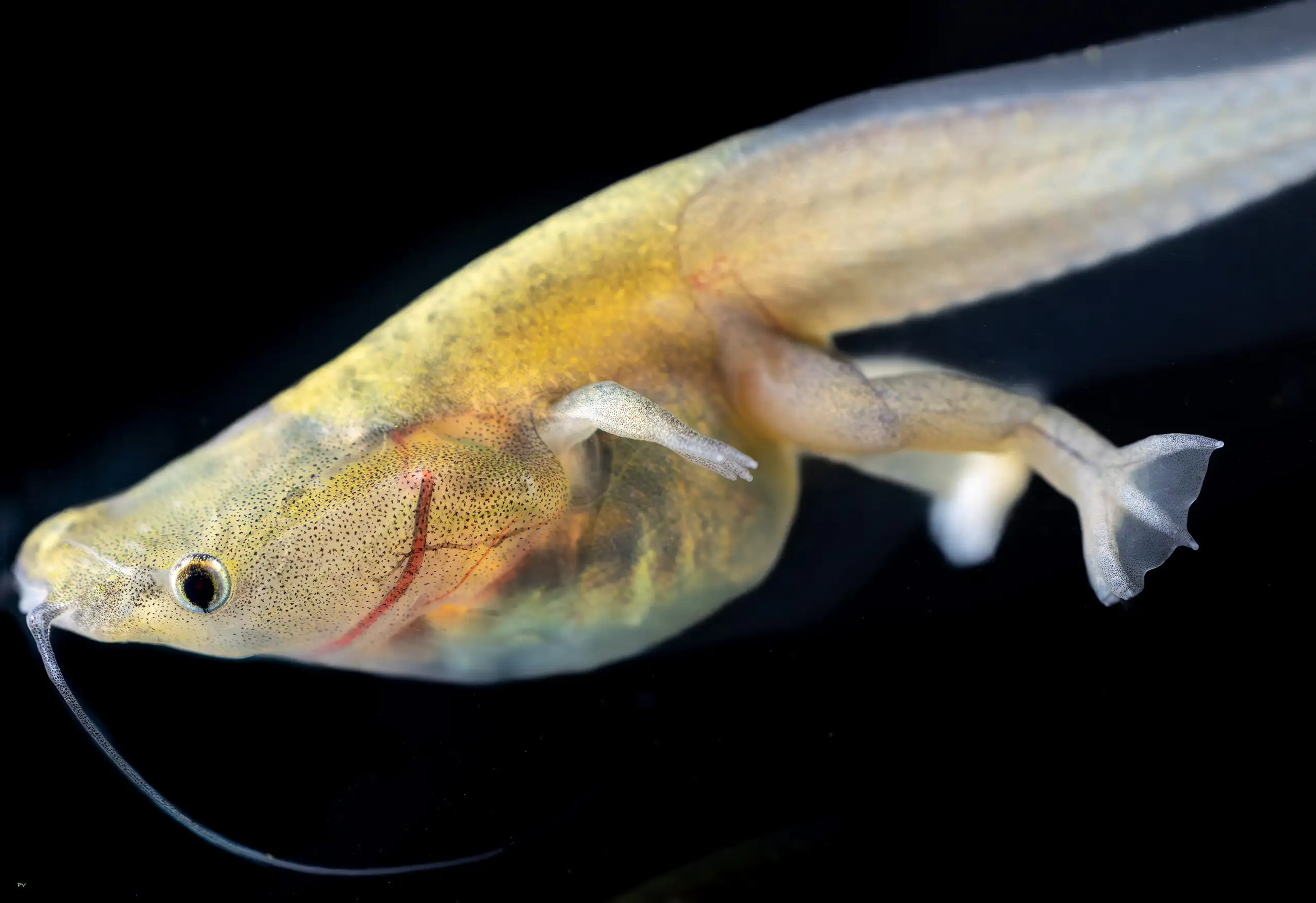

Why are frogs such a valuable animal model?
If you think about how valuable human twin studies have been in understanding human development, then consider what can be learned when you can study thousands of siblings at once. We know that the developmental basis of many diseases begins in the embryonic stage. But in humans we cannot easily see the development because the embryo is inside the mother. But with frogs you can see the embryos—tadpoles—developing in real time. You can manipulate the conditions again and again, allowing you to do large-scale experiments in a short period of time. The tadpoles are clear, right? So you can see some mutations as they occur. You can directly manipulate multiple genes in multiple ways and get results in three days. In mice, similar experiments would take months and months.
With such rapid advances in gene editing and organoid development, aren’t Xenopus and other animal model systems becoming less relevant?
Not really. Genome editing works very well in frogs, but you can do it in hundreds of embryos in a single experiment. Moreover, we still use Xenopus experiments to optimize approaches that we translate to human organoid protocols. It’s important for people to know that there’s still important research being done with these non-mammalian model systems. There are thousands of labs around the world studying everything from neural developmental disease to the gastrointestinal system, cell cycles, models of cancer, and so on. Without resources like Xenbase, much of that research would grind to a halt because people would struggle to retrieve the most simple information about the genes they are studying.
While it is true that organoids are getting closer and closer to fully mimicking human organ function, we’re still not there. And the process of making organoids is complicated enough and expensive enough that there’s still real value in doing certain forms basic research in Xenopus and other animal model systems.
How are you using Xenopus in research at Cincinnati Children’s?
We maintain a colony of about 1000 frogs and we use them extensively. Before attempting to study a gene variant of interest using human stem cells, we start with hundreds of variants in animal models. This allows us to determine which gene variants make sense to test further.
Right now, we are working on a large collaborative project led by my colleague Jim Wells (the other co-director of CuSTOM) involving 12 research centers around the world to study trachea-esophageal disorders such as tracheoesophageal fistula and esophageal atresia. This year, we used Xenopus and CRISPR genome editing to screen genetic variants identified from whole-genome sequencing of these patients. With this approach, we were able to discover the likely genetic causes of esophageal atresia, a life-threatening birth defect that we treat in our area of digestive clinic. { https://pubmed.ncbi.nlm.nih.gov/40412385/ } Now we have identified key risk genes that we are further studying in a human organoid model.
What’s next for Xenbase?
In 2022, Xenbase joined the Alliance of Genome Resources, a consortium funded by the NIH that brings seven model organism databases and the Gene Ontology Consortium together under one umbrella. The goal is to provide an integrated view of all this genomic data to biologists, clinicians, students and others.
The Xenbase platform has managed to continue across 25 years of so much technological change and we are continuing to advance. Like many others, we are exploring how to use artificial intelligence to accelerate bio-curation and support interpretation, so we can synthesize disparate data into knowledge. We recently got an outstanding score on our renewal application at the NIH – our fingers are crossed for another five years of funding.
I’m proud that Cincinnati Children’s has been facilitating Xenopus research worldwide for all these years. I think Xenbase will be useful for quite some time.
Don’t Miss a Post:
- Subscribe to the Research Horizons Newsletter
- Follow Cincinnati Children’s Research Foundation on Bluesky, X and LinkedIn
| Original title: | Xenbase: 25 years of integrating molecular and biomedical data from Xenopus. |
| Published in: | Genetics |
| Publish date: | Nov. 3, 2025 |
Research By

Our goal to elucidate the molecular mechanisms controlling the embryonic development of digestive and respiratory organs. We use a combination of Xenopus and mouse animal models, human pluripotent stem cells and cutting-edge genomics to investigate the underlying gene regulatory networks of organogenesis.