Scientists Discover a Peanut Allergy ‘Fingerprint’ That May Help Push Emerging Treatment to New Levels
Research By: Krishna Roskin, PhD | Scott Boyd, MD, PhD
Post Date: March 9, 2020 | Publish Date: March 5, 2020
About 1 million families in the United States live with the fear that their child might break into hives or feel their throats begin to swell shut from severe allergic reactions to peanut exposure.
While the U.S. Food and Drug Administration approved the first therapeutic, an oral immunotherapy, for the treatment of peanut allergy in January, scientists say much more research is needed to eliminate the threat of severe reactions from peanut allergies.
On March 26, 2020, Science will host a Facebook Live event focused on Hoh et al.’s work and on peanut allergy more broadly. Reporters and the public are welcome. Event to be hosted here: https:/
/ www. facebook. com/ ScienceMagazine
That work took a significant step forward with new findings published online March 5, 2020, in Science Immunology by researchers at Stanford University, Cincinnati Children’s, and Johns Hopkins University School of Medicine.
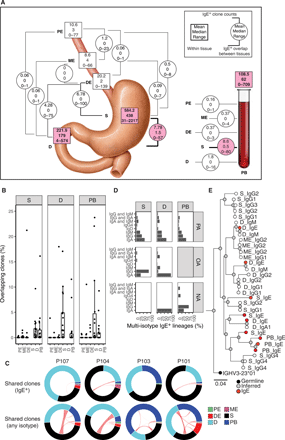
The research team collected tissue samples from 19 patients with peanut allergies, then conducted a series of advanced laboratory and bioinformatic analyses to determine which tissues house the immune cells producing the hyper-sensitive antibodies that trigger severe allergic reactions.
The team reports that patients with peanut allergies have a distinctly high concentration of immune cells located in stomach and duodenum, the small intestine adjacent to the stomach, which produce immunoglobulin E (IgE), the antibody type known to be responsible for severe allergic reactions.
This “fingerprint” of peanut allergy sensitivity was not seen as clearly in other locations, such as the esophagus or circulating in the blood. Furthermore, highly similar IgE antibody sequences were identified in multiple individuals with peanut allergy.
“Together, these findings indicate that recognizable and common IgE antibody gene rearrangements are a hallmark of peanut allergy,” the co-authors state. “Such sequence patterns, if replicated and extended beyond our relatively small clinical study, could begin to define a diagnostic set of IgE antibody types that may be specific for particular allergies and could be of value in tracking patient responses to treatment.”
The study was led by first author Ramona Hoh, PhD, and senior author Scott Boyd, MD, PhD, at Stanford.
Krishna Roskin, PhD, a faculty member in the Divisions of Biomedical Informatics and Immunobiology at Cincinnati Children’s designed and created the computational analysis platform used to analyze the antibody sequences. This included software to identify and track clonally related B cells across tissues from next-generation sequencing data.
Read the study in Science Immunology
| Original title: | Origins and clonal convergence of gastrointestinal IgE+ B cells in human peanut allergy |
| Published in: | Science Immunology |
| Publish date: | March 5, 2020 |
Research By








