‘Massively Parallel’ Gene Screening Tool Can Accelerate Research for Nearly Any Disease
Post Date: March 12, 2021 | Publish Date: March 12, 2021
Transformative discovery led by experts at Cincinnati Children’s began with a deep dive into lupus genetic mechanisms that led to an innovation that could boost studies of heart disease, diabetes, asthma and more
Ever since the discovery of DNA, scientists have raced to understand which tiny variations among the 3 billion base pairs within the human genome are responsible for causing disease.
While many are familiar with single-gene diseases such as cystic fibrosis or sickle cell anemia, science has lacked the computer and people power to sift through vast and expanding clouds of genetic data to tease out the patterns involved in complex conditions such as heart disease or asthma.
Now, as part of an ongoing exploration of systemic lupus erythematosus (SLE), a 16-member research team led by experts at Cincinnati Children’s reports a far-reaching discovery. They have developed a new variation on a “massively parallel” genetic screening tool that sheds new light on lupus—and potentially many, many more conditions.


“This study not only provides several critical new discoveries about lupus, it also provides a blueprint for dissecting the genetic mechanisms of many complex human diseases,” says Leah Kottyan, PhD, interim director of the Center for Autoimmune Genomics and Etiology (CAGE) at Cincinnati Children’s, who serves as co-senior author for this study along with Matt Weirauch, PhD.
Detailed findings were published online March 12, 2021, in Nature Communications.
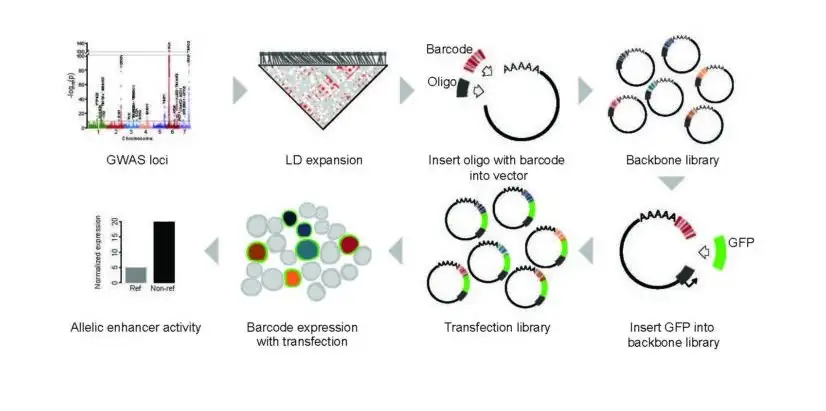
MPRAs help scientists quantify the ‘regulatory capacity’ of thousands of pieces of DNA in a given cell type. Such DNA sequences control how much, when, and where particular gene products are made.
“More than 95% of the genome does not code for genes, but instead is involved in other processes, such as controlling gene expression levels,” Weirauch says. “Understanding more about this information is vital because these regions of the genome also happen to be where genetic risk concentrates for complex diseases such as lupus.”
In this study, the authors use MPRAs to examine the effect of every known lupus genetic variant on gene expression levels.
Having this level of data available to researchers could open many doors for developing highly targeted therapies—and make clinical trial results much more informative, Kottyan says. For example, this kind of data could make it easier to approve a drug’s use for the people most likely to benefit, while precisely detecting those who should not take the drug at all.
Further down the road, this type of technology will be needed to bring a long-term hope for the future of medicine closer to reality: to become able to custom produce medications to the specific needs of individual patients.
LESSONS FROM CHASING LUPUS
Lupus is a painful disease that plays havoc with the body’s immune system. Up to 300,000 people in the US have lupus, and the disease causes or contributes to as many as 2,000 deaths a year, according to the US Centers for Disease Control and Prevention.
About 90% of those affected are women. Common symptoms such as fatigue, joint pain, fever and rash typically begin between ages 15 and 44. The disease can also damage the kidneys, brain, heart, and lungs.
Over the years, studies have documented numerous potential gene variations involved with lupus. A number of those early gene discoveries were made by researchers at Cincinnati Children’s from a lab led by John Harley, MD, PhD.
In 2018, a team led by Weirauch, Kottyan, and Harley reported that lupus was one of seven common chronic diseases that appear strongly linked to prior infections with the Epstein-Barr virus (EBV)—best known for causing mononucleosis. The team found that transcriptional regulators from the virus can greatly influence human gene expression levels, especially within the immune system. The increased disease risk that results for infected people varies according to which part of the genetic code was affected by the invader.
In several ways, the new study in Nature Communications expands upon that 2018 work.
For women battling lupus, this study boiled down more than 3,000 potentially important genetic variants to just 51, many of which appear related to B cell function, a core component of our immune systems.
The new genetic information provides immediate new ideas that could lead to more effective ways to prevent lupus-related complications, Kottyan says.
WHY IS A NEW GENETIC SCREENING APPROACH IMPORTANT?
Longer-term, the successful development of massively parallel gene screening has potential application for finding the genetic connections to virtually any disease. Here’s how:
For nearly two decades, scientists have been using genome-wide association studies (GWAS) to uncover large sets of clues about gene variations that might be causing a disease. The process compares entire genomes (3 billion base pairs) between people with a disease and others without that disease. The results include long lists of single-nucleotide polymorphisms (SNPs) that differ between the healthy and unhealthy people—only some of which are relevant to the disease being studied.
GWAS basically converts the proverbial hunt for a needle in a haystack into hunting for the right needle in a large box of interesting needles collected from many haystacks. Hunting through such boxes has produced major discoveries for a growing list of diseases, from Huntington’s disease to the BRCA-1 gene that increases breast cancer risk.
But some of the boxes of needles are huge. In conditions such as heart disease, diabetes or asthma, dozens or even hundreds of gene variations may be involved, all interacting with each other in exceeding complex patterns. In these situations, screening every needle in the box would take years.
Since GWAS emerged, researchers have devoted years to developing shortcuts that can help sort through the noise. For example, scientists often use databases that use the latest assumptions about likely areas of genetic activity to point investigators towards the most promising corner of the box of needles.
The major limitation: many studies continue to find relevant SNPs in unexpected places. The potential impact of MPRAs: Scientists now have an extremely fast tool for evaluating all the needles in the box.
“The sheer scale of the data that can be analyzed is astonishing,” Kottyan says. “For example, in 2019, I published a paper based on one reporter assay experiment. This paper includes the results of over 150,000 reporter assay experiments.”
Before using this method, it often took at least two years for scientists to understand the molecular mechanisms at work in a single genetic locus (or region of the genome) that may contain disease-causing SNPs. This study also took more than two years—but the team was able to analyze all 91 genetic loci known to increase lupus risk.
Crucially, the same screening process can be used to sift suspected genetic associations for nearly any complex disease.
“This is a huge acceleration of discovery,” Kottyan says.
NEXT STEPS
A flurry of related lupus research is beginning to explore the new clues revealed by the MPRA method.
“In terms of new treatment, we are using what we learned from this study to identify medications that would affect many lupus risk SNPs. Because each SNP increases risk by a modest amount, targeting many at the same time will likely have a higher value for patients,” Kottyan says.
The Cincinnati Children’s team also has begun using the MPRA technology to further explore atopic dermatitis, eosinophilic esophagitis, and multiple sclerosis.
ABOUT THE STUDY
Cincinnati Children’s co-authors in this study include 13 researchers from CAGE and the divisions of Biomedical Informatics, Immunology, and Developmental Biology. They include co-corresponding author Matthew Weirauch; co-first authors Xiaoming Lu and Xiaoting Chen; and co-authors John Harley, Emily Miraldi, Carmy Forney, Omer Donmez, Daniel Miller, Sreeja Parameswaran, Yongbo Huang, Mario Pujato, and Tareian Cazares.
Other collaborators include John Ray and Carl G. de Boer from the Broad Institute of Massachusetts Institute of Technology (MIT) and Harvard University; and Ted Hong, Department of Pharmacology and Systems Physiology at the University of Cincinnati College of Medicine.
Funding sources include grants from the National Institutes of Health (F32 AI129249, K99-HG009920, P30 AR070549, P30 DK078392, R01 AI024717, R01 AI148276, R01 AR073228, R01 DK107502, R01 GM055479, R01 HG010730, R01 NS099068, U01 AI130830, U01 HG008666); Cincinnati Children’s Hospital Research Foundation (Academic and Research Committee award, Center for Pediatric Genomics pilot funding, Endowed Scholar award); the Veterans Administration (I01 BX001834); the Ohio Supercomputing Center; and the state of Ohio.
| Original title: | Global discovery of lupus genetic risk variant allelic enhancer activity |
| Published in: | Nature Communications |
| Publish date: | March 12, 2021 |