Neonatal Antibiotics Disrupt Esophageal Microbiota, May Increase Risk of EoE
Research By: Michael Brusilovsky, MMedSc, PhD | Marc Rothenberg, MD, PhD
Post Date: October 13, 2021 | Publish Date: Oct. 7, 2021
When newborns need antibiotics to stave off potentially lethal infections, some of those who survive the short-term crisis may become more likely to experience a long-term problem—a higher risk of developing the severe food allergy eosinophilic esophagitis (EoE).
That’s one of the key findings from a study published Oct. 7 in Gastroenterology by scientists at Cincinnati Children’s, the University of Pittsburgh and the University of Chicago. The team performed a series of experiments involving mouse models to detail the roles esophageal bacteria play in driving food allergy.
The study is one of the latest in a growing body of evidence illustrating how human health depends in many ways upon other organisms living within us.
“The esophagus has a unique microbiome that interacts with the developing host immune system. Disruptions to that balance, if unaddressed, may pose significant implications for long-term health,” says corresponding author Marc Rothenberg, MD, PhD, Director of the Division of Allergy and Immunology at Cincinnati Children’s.
The investigators were able to manipulate the esophageal microbiome by transplanting fecal matter or using antibiotics.
“One of the remarkable findings in the study was that the stool could replenish the normal microbiome of the esophagus, indicating that the local environment shapes the bacterial content in each gastrointestinal segment,” says the study’s lead author, Michael Brusilovsky, MMedSc, PhD.
The study involved conducting 16S rRNA gene sequencing to identify all the commensal bacteria strains found in the esophageal epithelium of the mouse during the first months of life. The team then compared the data to a library of genes and signaling pathways associated with human EoE.
The team further demonstrated that antibiotic treatment can disrupt the microbial balance, and showed that human esophageal microbiota in EoE patients undergoes similar perturbations, which in turn can alter the developing immune system in ways that allow EoE to develop.
This finding in the esophagus adds to a large body of research about the importance of the microbiome in the development of allergy. Among those findings, the University of Chicago’s Cathryn Nagler, PhD, (a co-author on the new study) was the senior author of a study in PNAS in 2014 that found that commensal bacteria microbiota regulates uptake of dietary antigens in the gastrointestinal tract and helps to protect the body from food allergen sensitizations.
Looking ahead, the research team plans to examine the genetic interaction with the microbiome changes and determine the feasibility of translating the findings for preventive medicine and therapies.
Read More About EoE Research at the Rothenberg CURED Lab
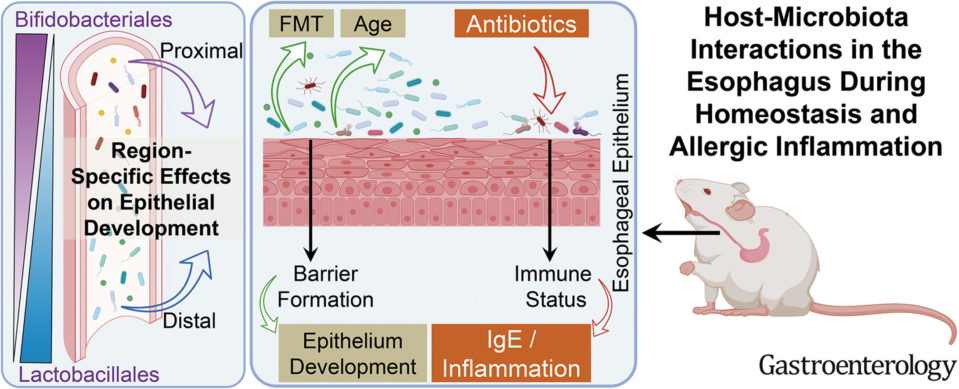
| Original title: | Host-microbiota interactions in the esophagus during homeostasis and allergic inflammation |
| Published in: | Gastroenterology |
| Publish date: | Oct. 7, 2021 |
Research By