Two Discoveries Boost Next-Generation Organoid Development
Research By: Aaron Zorn, PhD
Post Date: August 27, 2020 | Publish Date: Aug. 27, 2020
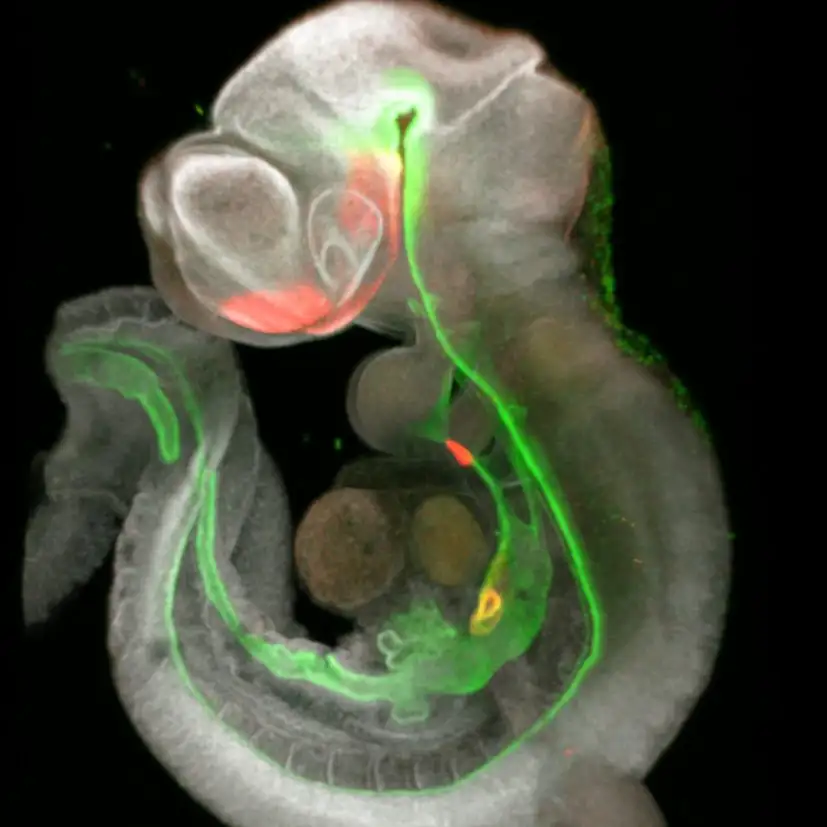
New research collaboration tunes into signals that will help make better organoids to mimic functions of the trachea, esophagus, stomach, liver, gallbladder, bile ducts and pancreas
In back-to-back reports published Aug. 27, 2020, in Nature Communications, a team of scientists from Cincinnati Children’s and Japan report discoveries that will be vital to a new wave of more-complex organoid development.
Their findings advance efforts to use human stem cells to grow organs from the fetal foregut including the trachea, esophagus, stomach, liver, gallbladder, bile ducts and pancreas.
“With single-cell analysis of mouse embryos, we defined the complex signaling networks controlling development of mesenchyme cells, which form the smooth muscle and fibroblast tissues that are essential for organ function,” says senior author Aaron Zorn, PhD, who leads organoid development at Cincinnati Children’s. “We then used this information from the mouse to differentiate the equivalent human tissue in the lab. This is important because up until now all of the liver, lung, stomach and esophagus organoids that we make mostly lack these mesenchyme cell types.”
Zorn directs the Center for Stem Cell & Organoid Medicine (CuSTOM) at Cincinnati Children’s, which has made groundbreaking advances in stomach, intestine, liver and esophageal organoid development. In 2019, the CuSTOM group launched a formal collaboration with RIKEN, Japan’s largest comprehensive research institution, to pursue further organoid innovation.
The papers published in Nature Communications represent the first results of that collaboration.
Decoding foregut development cell-by-cell
In this study, the scientists report detecting a set of signals within the foregut—a proto-organ in very early-stage embryos—that trigger how and when the other organs form. Specifically, they found that the signals are driven by the genes Wnt and SHH, which travel between cells in the endoderm and mesoderm layers of very early embryos.
To define these signals, co-first authors Lu Han, PhD, and Keishi Kishimoto, PhD, collaborated with organoid experts James Wells, PhD, and Takanori Takebe, MD, to develop a high-resolution map of foregut development in mice. They detected an unexpected variety of cells sending a chorus of master signals that trigger formation of the various organs that branch out from the foregut.
This study is the first to pin down the dynamics at play in the embryonic mesoderm, co-authors say.
The action happens very early–between embryonic days 8.5 and 9.5 in mice, which roughly corresponds to days 17 to 23 in human gestation. During this brief window of development, groups of cells at certain spots along the simple foregut tube begin transforming into the sprouts of organs that become the trachea, esophagus, liver and pancreas.
By studying the molecular signaling activity during this period, at a cell-by-cell level, the researchers produced a roadmap that shows how and why the organs sprout where they do. They then used these signals to grow tissue into different organs from human pluripotent stem cells.
In September 2019, Takebe and colleagues reported the world’s first success at growing a three-organoid system that included the liver, pancreas and biliary ducts. That breakthrough took five years to achieve and the organoids produced did not possess all the cell types needed for full-sized function.
The new road map will enable CuSTOM scientists to grow more compete interconnected organs, Zorn says.
A deep dive into trachea development
In a parallel paper also appearing in Nature Communications, the RIKEN and CuSTOM teams extended these studies with extensive experiments in mice to further define the mechanisms of trachea formation.
This study led by lung development expert Mitsuru Morimoto, PhD, in Japan used genetically modified mice to learn which cell signals were most important to trachea formation. When these signals fail, the developing embryo does not properly form the cartilage rings and smooth muscle tissues that the trachea needs to pipe air to the lungs.
Cincinnati Children’s, which developed the first human esophagus organoid in 2018 has been working with the RIKEN team on this project as part of its involvement with the CLEAR Consortium (Congenital Esophageal and Airway Defect Research).
“This work helps explain what happens when birth defects like esophageal atresia, tracheoesophageal fistula, and tracheomalacia occur,” Zorn says. “This work also opens the door to one day generating esophagus and trachea tissue for tissue replacement.”
Implications for tissue engineering
The sheer complexity of the new signaling roadmap helps explain why it took so long to make the initial three-organoid breakthrough. For example, the map revealed five distinct populations of mesenchymal cells involved in liver formation alone.
Now, co-authors say the new roadmap will make the process faster, could expand the types of organs that can be grown together, and will allow researchers to grow sets of organoids engineered to mimic conditions that lead to birth defects or increased disease risk—including some forms of cancer.
“One important outcome of our study was to use the signaling roadmap to direct the development of stem cells into different organ cell types,” Takebe says. “This approach may have important applications for tissue engineering.”
Short term, such organoid systems can be used to test new medications with far less dependence on animal models, or to evaluate the harms caused by pollution, unhealthy diets, allergens, and so on. Longer term, once expects learn ways to grow organoids to significantly larger sizes, customized lab-grown tissues could be used to repair damaged organs and someday even replace failing ones.
The paper lays out the protocols for other scientists to use to make their own organoid systems. Detailed data collected during the project also can be explored via the interactive website https://research.cchmc.org/ZornLab-singlecell.
| Original title: | Single cell transcriptomics reveals a signaling roadmap coordinating endoderm and mesoderm diversification during foregut organogenesis |
| Published in: | Nature Communications |
| Publish date: | Aug. 27, 2020 |
Research By