Pain-Free Tape Strips Produce Powerful Genetic Data for Asthma and Allergy Research
Research By: Gurjit Khurana Hershey, MD, PhD
Post Date: April 20, 2020 | Publish Date: April 19, 2020
When it comes to studying the causes, risks and treatments for atopic dermatitis, scientists say beautiful data really does run skin-deep.
In findings published online April 19, 2020, a team of Cincinnati Children’s investigators led by Gurjit Khurana Hershey, MD, PhD, reports that the cells obtained from tape strip skin sampling yield a wealth of data that many had believed possible only from more-invasive skin biopsies. Details appear in the Journal of Allergy and Clinical Immunology (JACI).
“Our data demonstrate that this methodology simultaneously samples the surface skin biome as well as the underlying human DNA and RNA, which can be used to characterize host responses to environmental cutaneous exposures,” the co-authors write.
“This novel methodology is unique in that it utilizes dissolvable tapes and each single tape strip yields sufficient nucleic acids for metagenomics (biome characterization), keratinocyte epigenetic studies, and RNA expression studies. This is highly relevant as the biome varies by depth of sampling and this method enables characterization of each layer sampled by an individual tape.”
Not only did the team report success at collecting desired information at varying layers of skin, they evaluated several brands and types of tape. One type—a water-dissolvable tape designed for hydrographic applications—emerged as the most-consistent performer.
The team tested the tape strip method with 400 children (ages 1 and 2) who had been diagnosed with atopic dermatitis. The non-invasive method proved to be safe, well tolerated, efficient, and low-cost.
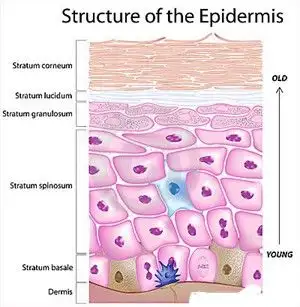
The tape strip method involved collecting skin cells by applying up to 11 strips, one at a time, to the same location. Each strip pulls cells from deeper layers, going from the mostly dead cell layers of the stratum corneum down into the living (thus fundamentally different) cells of the epidermis.
Near the surface, the first few tape strips revealed an expected variation in the distribution of microbial species. This finding was important because previous research has shown that the depth of S. aureus penetration into the dermis of atopic dermatitis lesion sites is associated with the presence of increased inflammatory cytokines.
At deeper layers, the tape strips pulled cells with enough consistency to show depth-based variations in gene expression that can help scientists understand the mechanisms involved with atopic dermatitis and why it so often leads to increased risk of severe asthma.
Previously, this kind of data could be obtained consistently only by skin biopsy, which is more painful, higher cost and less practical for large-cohort studies.
Given the practicality of the testing method, the co-authors say tape strip testing will have broad potential for research, especially for clinical studies.
In addition to Hershey, Cincinnati Children’s co-authors included: Mariana Stevens, PhD, Tammy Gonzalez, BS, Eric Schauberger, DO, PhD, Asel Baatyrbek kyzy, BSb, Heidi Andersen, MD, Daniel Spagna, BS, Mehak Kalra, BA, Lisa Martin, PhD, David Haslam, MD, Andrew Herr, PhD, and Jocelyn Biagini Myers, PhD.
| Original title: | Simultaneous skin biome and keratinocyte genomic capture reveals microbiome differences by depth of sampling |
| Published in: | Journal of Allergy and Clinical Immunology |
| Publish date: | April 19, 2020 |
Research By