Bone Marrow ‘Map’ Opens Path to Organoid-like Blood Stem Cell Production
Research By: Daniel Lucas, PhD
Post Date: February 10, 2021 | Publish Date: Feb. 10, 2021
“We finally have the tools to directly observe bone marrow cell differentiation.”
Daniel Lucas, PhD
Imagine a day when clinicians treating people with blood diseases such as leukemia or multiple myeloma can send in requests for laboratories to custom-produce specific types of blood cells to replace those affected by the disease.
That day became one step closer to reality with a new study led by experts at Cincinnati Children’s that provides powerful new insights into how bone marrow tissue works.
The study, published Feb. 10, 2021 in Nature, was led by senior author Daniel Lucas, PhD, and first authors Jizhou Zhang, MD, and Qingqing Wu, PhD, from the Division of Experimental Hematology and Cancer Biology. Co-authors include a team of scientists from Cincinnati Children’s and the University of Cincinnati, plus collaborators in Colorado, Texas and Michigan.
The team used a combination of cell-by-cell analysis techniques to build the first “atlas” of bone marrow tissue. The findings advance scientific understanding of how tiny blood vessels organize the bone marrow and regulate how blood gets produced.
“We finally have the tools to directly observe bone marrow cell differentiation. These results show that the bone marrow is a highly organized tissue and that this organization is provided by specific subsets of vessels,” Lucas says. “This is telling us that the organization of the vasculature dictates blood production. If we determine how the vessels function we will be one step closer to controlling the production of specific blood cells at will.”

A blueprint of blood cell production in situ
With recent advances in genomic analysis technology, researchers at Cincinnati Children’s and other major centers have been producing a growing library of “maps” that describe human tissue development in unprecedented detail. (Read more about the multicenter Pediatric Cell Atlas project.)
This study in Nature adds a thick reference book of bone marrow and blood cell development information to the virtual shelves.
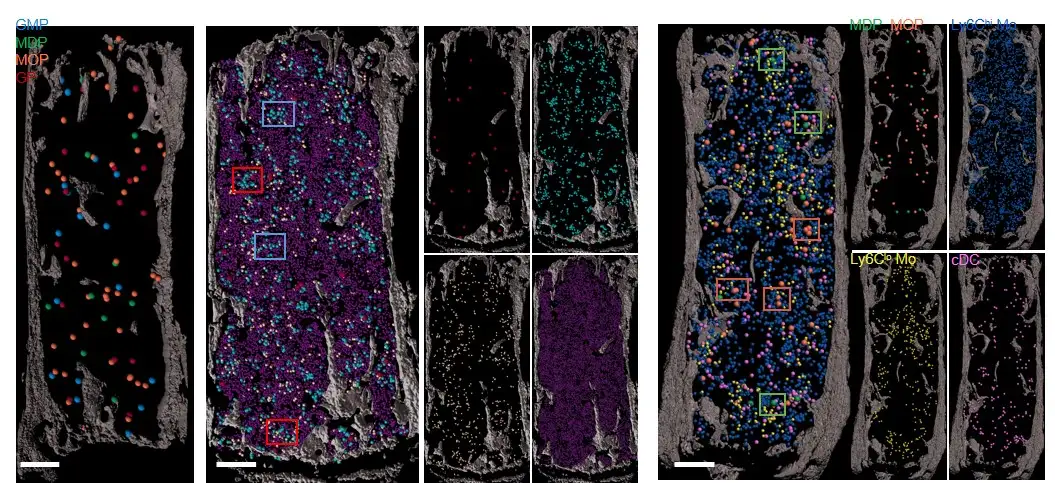
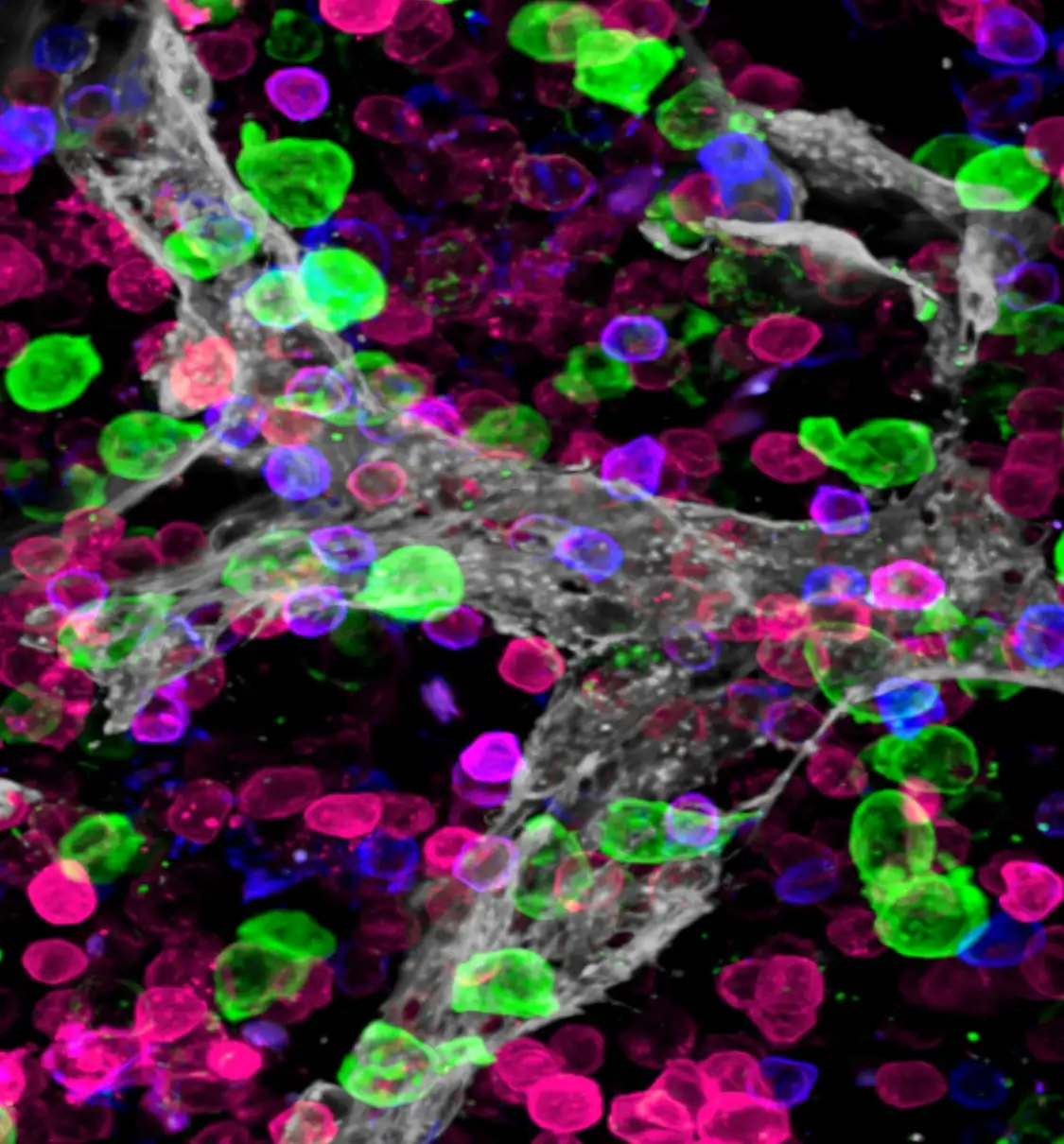
Making this bone marrow atlas required the development of novel confocal imaging methods for unprecedented resolution of blood cells in the marrow.
Until now, tracing the lineage of cell types during stages of development required destroying the tissue. In this project the team developed ways to image and trace for the first-time unique progenitors within the larger mass of bone marrow cells—without destroying the tissue structure.
“Tracking these special cell clusters revealed new information about the structure of bone marrow,” Lucas says, “including the insight that the bone marrow has a surprising degree of organization and that specific blood vessels support the production of unique blood cell types.”
How can this discovery advance care?
It typically takes several years of further research to translate basic science discoveries into practical applications. One potential direction for this line of study could be to support future development of highly customized blood cell factories that would mimic bone marrow function in laboratory settings.
Such blood organoids could be used to produce populations of blood cells with specific genetic variations, which scientists could analyze to develop improved treatments for disease. For example, a controlled method of blood cell production would be useful to scientists studying the immune system and how bodies defend themselves against infection, Lucas says.
Eventually, blood organoids might become a form of treatment themselves by allowing clinicians to replace a patient’s diseased cells with gene-edited healthy cells that face no risk of rejection. Having new understanding of how blood cells are produced in the bone marrow will help this effort.
“This certainly has implications for generating blood organoids,” Lucas says. “The groups working on blood organoids have been trying to produce organoids that can maintain or expand stem cell production. Our data indicates that additional structures are needed to produce mature blood cells in a balanced manner.”
Next steps
More study is needed to identify the molecular signals produced by the different blood vessels that regulate production of each type of blood cell. Knowing those signals will be a crucial step to regulating blood cell production.
Meanwhile, the research team plans to continue its work developing tools to image blood cell production, especially to expand imaging to earlier-stage progenitors.
“We want to be able to visualize every step in the production of the different types of blood cells,” Lucas says.
About this research
Funding for this study included grants from the National Heart Lung and Blood Institute (R01HL122661, R01HL136529); NIH/NCATS (U2CTR002818, NIH/NHLBI U24HL148865), and NIH/NIAID (U01AI150748); other NIH grants (R01AI120202, R01AI124657, DP1AI131080, AG045040, T32 AI118697, S10OD023410); and the US Department of Defense (W81XWH-20-1-0870 #CA191188).
Other funders include the Cincinnati Pediatric Cell Atlas Center, the HHMI Faculty Scholar’s program, March of Dimes Ohio Collaborative for Prematurity Research, the Burroughs Wellcome Fund, and the Welch Foundation.
| Original title: | In situ mapping identifies distinct vascular niches for myelopoiesis |
| Published in: | Nature |
| Publish date: | Feb. 10, 2021 |
Research By